Introduction
Transfection is one of the most commonly used techniques in molecular biology (Kim et al. 2010), and is the process of introducing DNA or RNA into target cells, typically in a circular form known as a plasmid. DNA/RNA are hydrophilic and typically cannot pass through the hydrophobic cell membrane, necessitating the use of transfection techniques in order to internalize them and study gene function and protein expression within the cell.
Different transfection methods have different efficiencies and levels of overall gene expression, and may even affect cell viability (depending on the cell type used). In order to evaluate transfection efficiency, sensitive assays are required so that transfection can be optimized.
Transfection Methods
There are a range of transfection methods, resulting in transient or stable transfection:
- Transient: Transfected DNA remains in the cytoplasm, not incorporated into chromosomal DNA in nucleus. Cells will lose the transfected DNA after a few divisions.
- Stable: Transfected DNA incorporated into chromosomal DNA, persists in daughter cells upon division. Stable cell lines useful for repeated experiments.
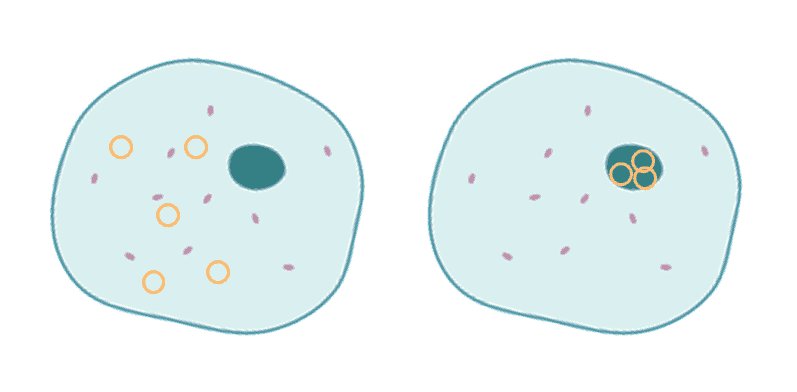

| Figure 1. Transient vs stable transfection. DNA plasmids (yellow circles) have been successfully transfected into a eukaryotic animal cell. The left is transient transfection, the plasmids remain in the cytoplasm. The right is stable transfection, the plasmids have entered the nucleus and become incorporated into the cell genome. |
Transfection methods can be grouped into biological, chemical or physical methods.
Biological methods use viruses and are therefore also known as ‘transduction’. While being highly efficient, viruses cannot carry much genetic material and insert the material randomly into the host genome, potentially causing inflammatory and cytotoxic reactions if critical housekeeping genes are disrupted.
Chemical methods use positively charged chemicals that are attracted to the negatively charged cell membrane and pass through via endocytosis. Popular chemicals used are polymers, calcium phosphate (CaPO4), lipids and amino acids. The most popular method uses lipids, which form around the target DNA in a lipid: DNA complex, which can pass into the cell.
Physical methods involve either directly injecting the DNA into the cell, or creating pores in the membrane for the DNA to pass through. The latter can be done with electricity (electroporation), lasers (phototransfection), sound (sonoporation) or even magnetism (magnetofection).
Transfection Efficiency
The ideal transfection method has high transfection efficiency, low cell toxicity, minimal secondary effects on the cell, easy to use, and reproducible. While some cell lines are optimized for transfection with robust protocols, use of multiple cell types will require optimization. Optimization typically involves growing cell in a multi-well plate with controls, and altering the concentration and ratio of transfection reagents across the plate. Different transfection methods and reagents can be tested, as well as incubation time, cell density, growth medium, etc.
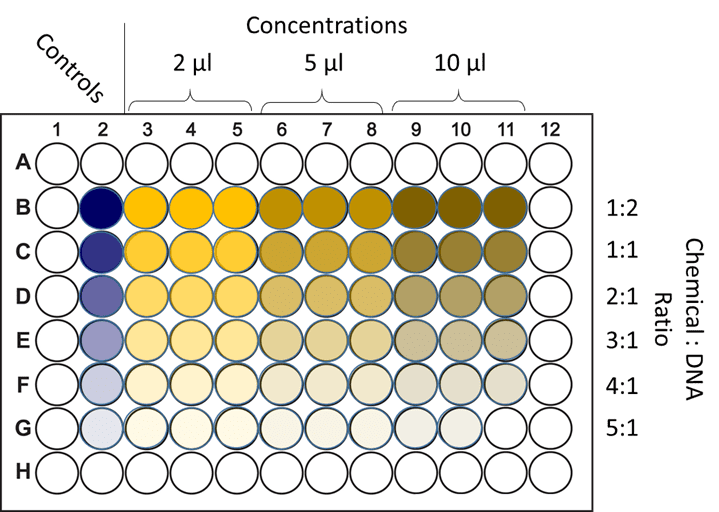
| Figure 2. Optimizing chemical transfection. Cells are grown across a multi-well plate and exposed to different transfection conditions. Either the concentration of DNA is altered (across the columns), or the ratio of the chemical factor (such as lipids) to DNA is altered (down the rows). These conditions are all then tested with a transfection efficiency assay to determine the optimum condition for this method and cell type. |
When transfection has occurred, transfection efficiency assays are used to evaluate the success of this transfection method. Cell DNA itself can be analyzed, or downstream consequences such as protein expression. Quantifiable methods are required to determine how successful transfection was, requiring sensitive markers or reporter genes (Sandbichler et al. 2013).
Plasmids typically include reporter genes that code for markers not found in the cell. If these genes are unique to the plasmid and the protein markers are expressed within the cell after transfection, it can be determined to be successful, the efficiency of transfection proportional to the signal intensity. These reporter gene assays can be performed several days after transfection but this is another factor to optimize.
Popular reporter gene markers used for these assays include:
- Fluorescent/luminescent markers such as GFP/luciferase, these can be easily detected using fluorescence microscopy techniques across a wide range of cells in a high-throughput and quantitative manner. With fluorescent markers flow cytometry is especially effective.
- Enzyme markers such as β-galactosidase, this is an coli enzyme that reacts with X-gal to stain cell blue, allowing for simple yet quantitative visual assessment.
- Media markers such as secreted alkaline phosphatase (SEAP), this marker is secreted from cells into the media, which can be taken from the cultures at various time for analysis. This method does not result in cell damage, unlike some enzymatic or luminescent methods.
While many assays use a single reporter gene, some use dual or multiple reporters in order to normalise expression against cell number and transfection efficiency which results in more robust results. Other examples include staining cells with multiple markers, using GFP-marked cells to quantify successful transfection, Hoechst/DAPI to determine the overall cell number, and a third stain to quantify cell death.
An alternative to reporter genes is to simply use assays that determine the expression levels of DNA/RNA/protein in the cells, this can be done with RT-qPCR or (for DNA or mRNA) or with ELISA/Western blots (for protein). If using a cell line that has consistent stable transfection, results can be compared to transiently transfected cells.
Summary
Transfection is a popular technique enabling researchers to investigate protein function and gene regulation, across different cell types in different physiological or pathological conditions. DNA/RNA is forced into cells via biological, chemical or physical methods, and the efficiency of transfection can be determined using a wide range of different assays, which rely on markers placed into the DNA/RNA, or the use of sequencing techniques to directly quantify transfection results.
References
Kim, T. K., & Eberwine, J. H. (2010). Mammalian cell transfection: the present and the future. Analytical and bioanalytical chemistry, 397(8), 3173–3178. https://doi.org/10.1007/s00216-010-3821-6
Sandbichler, A. M., Aschberger, T., & Pelster, B. (2013). A method to evaluate the efficiency of transfection reagents in an adherent zebrafish cell line. BioResearch open access, 2(1), 20–27. https://doi.org/10.1089/biores.2012.0287